Applying Deep Learning to graph analysis is an emerging field that is yielding promising results. Our approach WalkRNN described below leverages research in learning continuous feature representations for nodes in networks, layers in features captured in property graph attributes and labels, and uses Deep Learning language modeling to train the computer to read the ‘story’ of a graph. The computer can then translate this learned graph literacy into actionable knowledge such as graph classification tasks.
Graph’s flexibility presents challenges Graph provides a flexible data modeling and storage structure that can represent real-life data, which rarely fits neatly into a fixed structure (such as an image fixed size) or repeatable method of analysis. Graph heterogeneity, node local context, and role within a larger graph have in the past been difficult to express with repeatable analytical processes. Because of this challenge, graph applications historically were limited to presenting this information in small networks that a human can visually inspect and reason over its ‘story’ and meaning. This approach fails then to contemplate many sub-graphs in an automated fashion and limits the ability to conduct top-down analytics across the entire population of data in a timely manner. Deep Learning is an ideal tool to help mine graph of latent patterns and hidden knowledge.
In the past, it has proven difficult to apply machine learning algorithms to graph. Methods often reduce the degrees of freedom by fixing the structure in a repeatable pattern, such as looking at individual nodes and their immediate neighbors, so the data can then be consumed by tensor-oriented algorithms. The rich information captured by the local graph topology can be lost with simplifications, making it difficult to derive local sub-structures, latent communities and larger structural concepts in the graph.
Research — Machine Learning applied to Graph Research in the past few years has made strides in a class of approaches that learn, in an unsupervised way, continuous feature representations for nodes in networks, such that features are sensitive to the local neighborhood of the node. With these feature representations (stored in vector space), nodes can be analyzed in terms of the communities they belong to or the structural roles of nodes in the network. Jure Leskovec and others at Stanford have contributed research and performant algorithms in this space:
-
The node2vec paper gives a background of research that uses the Random Walk method to represent a network as a ‘document’ that is sensitive to edge weight and local topology of the starting node (similar to the SkipGram Word2Vec algorithm).
-
GraphWave provides an effective way to capture the structural role of nodes, such that “nodes residing in different parts of a graph can have similar structural roles within their local network topology”.
Deep learning graph classification and other supervised machine learning tasks recently have proliferated in the area of Convolutional Neural Networks (CNNs). The DGCNN team (2018) developed an architecture for using the output of graph kernel node vectorization (using struct2vec, in a similar space as GraphWave) and producing a fixed sorting order of nodes to allow algorithms designed for images to run over unstructured graphs.
Our results below are compared to the DGCNN paper (and related benchmarks) to illustrate how a language model (RNN) can also be used to classify graphs. In cases where rich information is stored in graph properties (e.g. attributes and labels), our approach produces superior results and can potentially be applied to cases of free text passages stored in graph properties (look for future posts on this topic).
Language Model over graph Property graph can contain contextual and rich information in its properties, both on the nodes and edges, and also captures information about larger network organizations and how the parts interact to create the whole. The goal of our approach is to learn the ‘story’ of a sub-graph. We accomplish this through a few simple steps.
-
Our code WalkRNN first enriches an existing property graph with featurization such as the structural role of nodes(using GraphWave) and collapses categorical and continuous data in attributes and labels into repeated ‘words’ globally present within the graph.
-
Then we use code from node2vec’s biased random walk to create ‘sentences’ of nodes and edges (an approach which the node2vec paper notes is computationally efficient in terms of both space and time requirements and is shown implemented in Apache Spark). The sentence of a random walk for a small graph of a family, enriched by WalkRNN, might read (if translated into human readable words): ‘Mary age_class_3 gender_F structural_hub_node edge_is_sibling Martha age_class_4 gender_F structural_leaf_node edge_is_sibling Lazarus age_class_4 gender_M structural_connector_node…’ and so on until the walk length is reached. Every node spawns multiple random walks. In the process sub-structures such as triangles can lead to loops or journey into new parts of the graph depending on the random decisions of the walker.
-
For our deep learning, we use the fast.ai library’s language model and text parsers and pass the ‘documents’ of random walks into an RNN (AWD_LSTM) which retains memory of past words in the sequence and learns meaning behind the ‘words’ and their relationships. In this way, the graph relationships between nodes and their local context are learned by the computer. Patterns held across the separate components are then learned as well, as the ‘words’, such as the node structural role or property categorical value, are often shared across components and begin to mean something in relation to other ‘words’.
-
For every component, our code randomly selects walks and concatenates them together into a ‘paragraph’ of cuts through the graph, ideally long enough to cover the scope of the graph regions and to capture the gist of the component type. Finally, these paragraphs per component are passed into an RNN classifier using transfer learning from the pre-trained RNN language model and predicts the label for the component.
WalkRNN vs DGCNN in graph classification The effectiveness of this method really shines for certain types of graphs (in the AIDS case, for example, it correctly predicted the class for 400 out of 400 graphs), and the results seem to speak for themselves.
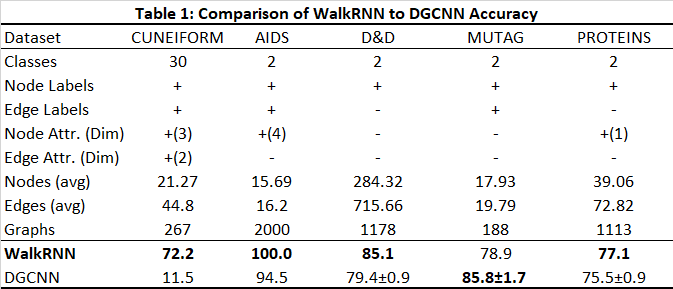
The results above are dependent on parameters (such as dropout, learning rate, neural network # hidden layers and #RNNS, walk length, # structural GraphWave ‘words’), and repeated runs were required to fine-tune results. The DD data set only includes node labels and edges (ie no node/ edge attributes or edge labels), and the power in enriching the graph ‘story’ with properties is not really demonstrated. MUTAG seemed less stable in training as there were so few examples (only 188 graphs).
The graph kernel datasets and accompanying stats used for these experiments were downloaded from this website provided by the TU Dortmund Dept of Computer Science. The datasets share a common format, making it easy to experiment with graph classification across multiple domains. Each dataset is broken into multiple graphs, each with its own class label. Our code learns how to read the various graph domains from scratch and then learns how to predict the class label for each graph (e.g. AIDS or not AIDS).
The accuracy in Table 1 shows our accuracy at predicting test graphs (typically 20% random sample of the entire set). The only exception was in the case of the larger DD dataset in which the training set was 12.8% of the set, validation 3.2% and testing 4%. The results for DGCNN were taken both from the creators’ paper and from this blog where the DGCNN tool was run on various datasets (such as Cuneiform and AIDS).
If you find this approach interesting or helpful, please feel free to email us at info@tylordata.com for further information.
Community Inspiration Special thanks to the DGCNN team, Jeremy Howard and his fast.ai deep learning project, Jure Leskovec and researchers at Stanford, and TU Dortmund Dept of Computer Science for their great contributions to the community of graph and deep learning enthusiasts.
WalkRNN is the brainchild of Deborah Tylor, owner of Tylor Data Services, LLC. Key contributor to the code and ideas is Joseph Hagaa. Thanks also to Mirco Mannucci and Elena Romanova for inspiration and Sook Seo for moral support and graphical aids. And, in everything, thank you Jesus.